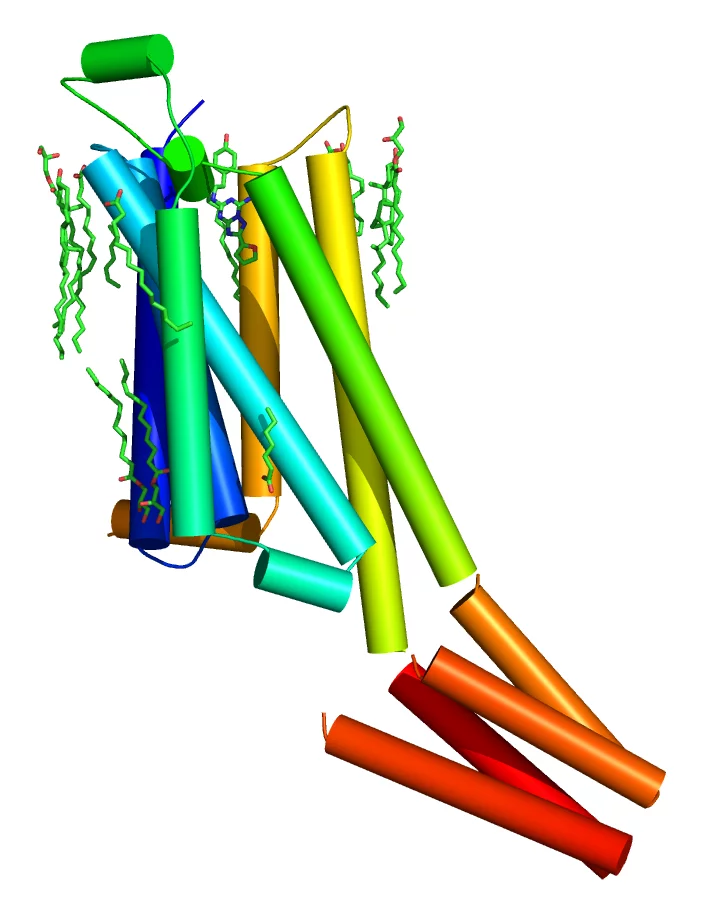
Serial femtosecond crystallography (SFX) experiments were performed using long-wavelength X-ray pulses (4.5 keV) from the SwissFEL accelerator, which increased the anomalous signal from naturally occurring heavy elements in proteins such as sulfur and phosphorus. This increased signal was employed in native-SAD phasing of SFX data for the first time and consequently reduced the sample consumption and beam time significantly as compared to other native-SAD phasing experiments at other XFELs. Even more remarkable was that these results came from the first ever protein crystallography experiment performed at SwissFEL. IUCrJ is an open-access publication and the paper can be downloaded here. The structure of the A2A membrane protein is shown below.
In addition to an increased anomalous signal at longer wavelengths, this significant reduction is attributed to advances in data analysis methods and the low-noise JUNGFRAU 16-mega pixel detector, which is the largest area detector operated at an XFEL to date. Improvements in data analysis presented in the article reduced the negative impact on data quality from partial intensities and inaccurate experimental geometry in SFX typically observed in SFX data sets. This significant achievement can facilitate de novo, radiation-damage-free structure determination of small and weakly diffracting protein crystals at XFELs, a potential that remains heavily underexploited especially by the pharmaceutical industry. This is particularly important for membrane proteins that do not crystallize readily to a size sufficient for synchrotron studies. Several other XFEL facilities worldwide are capable of reaching X-ray beam parameters including the tender X-ray energy regime (2–5 keV), providing excellent opportunities for long-wavelength native-SAD phasing of difficult targets.
For further information on this result please contact Dr. Karol Nass (+41 56 310 3978) who is the lead author of the paper. Further information on our pharmaceutical collaborators LeadXPro and Heptares can be found on their websites.