Your local contact will provide you with the latest version of our cxs_software package upon the start of your beamtime, or you can request them earlier from your local contact. These macros provide the functionality for reading and displaying the detector frames and for example for azimuthal integration. A zip-file with these macros can be downloaded here as well. The macros are provided as-is without any guarantees or liability on our part. An introduction to some of the packages is available in the cSAXS notes (see under manuals). Besides the basic codes below you can find other packages for example to analyze and visualize orientation and degree of orientation of scanning SAXS, reconstruct and visualize SAS tensor tomography, and ptychography reconstruction code. The codes are provided "as they are" and without any guarantee of results, performance, or support. The user of the codes should always take responsibility to ensure the correctness of the results.
Note: If these codes, or subfunctions or parts of it, is used for research in a publication or if it is fully or partially rewritten for another computing language the authors and institution should be acknowledged in written form in the publication. For example "Data processing was carried out using the "cSAXS matlab package" developed by the CXS group, Paul Scherrer Institut, Switzerland." Variations on this text can be incorporated, upon discussion with the CXS group, if needed to more specifically reflect the use of the package, or function, for the published work. A publication that focuses on describing features, or parameters, that are already existing in the code should be first discussed with the authors.
Data format - NeXus H5
Data and metadata is provided in one H5 file per scan. We organize the H5 file contents following NeXus guidelines and in particular following NXsas as much as it is practical.
The files can be explored and contents read with standard tools such as h5py. We also provide routines in Matlab to read it. In particular within the base package below we have the function io.nexus_read
% read the whole file contents
nxs = io.nexus_read('~/Data10/data/S00000-00999/S00250/e16812_1_00250.h5');
% let the function try to find the file, read whole file contents
nxs = io.nexus_read('~/Data10', 'scanNr', 250, 'filter', '/');
% load only spec data
nxs = io.nexus_read('~/Data10', 'scanNr', 250, 'filter', 'spec');
% load only pilatus_1 data
nxs = io.nexus_read('~/Data10', 'scanNr', 250, 'filter', 'pilatus_1');
% load all detectors
nxs = io.nexus_read('~/Data10', 'scanNr', 250, 'filter', 'detectors');
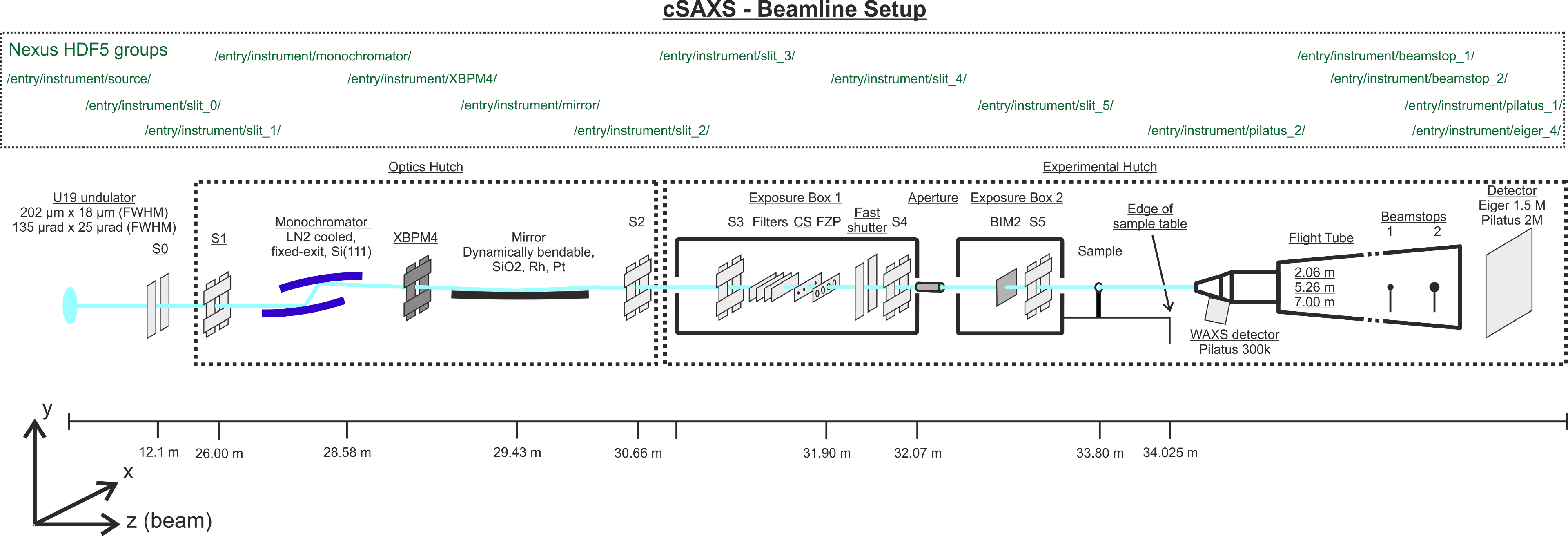
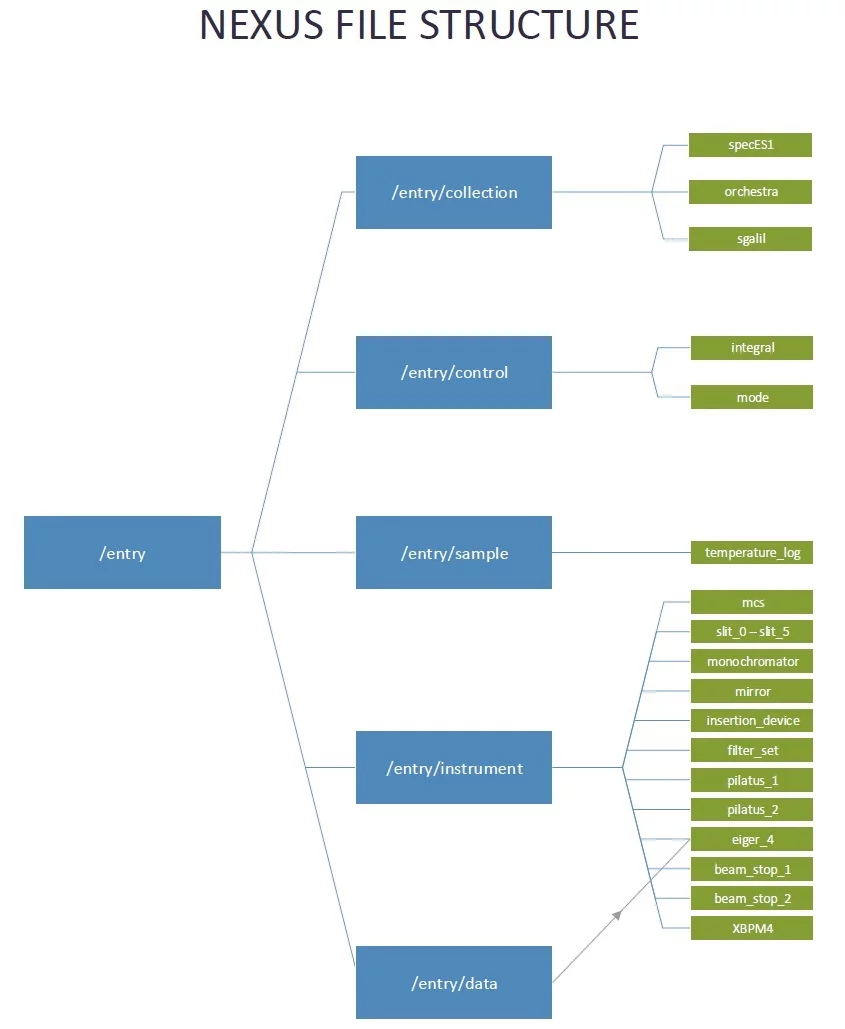
The following show 1) a sketch with beamline components and the corresponding H5 groups, and 2) the organization of the H5 groups.
cSAXS beamline software packages
Base package
Basic functionalities for file reading and radial integration and plotting. Also provides a lot of functions used in the other packages.
Scanning SAXS package
Analysis and plotting of scanning SAXS, main orientation of scattering, degree of orientation. Please cite Bunk et. al. New J. Phys. 11, 123016 (2009). Uses the base package above.
SASTT package
Reconstruction of small-angle scattering tensor tomography (SASTT), which provides information about 3D anisotropic nanostructure within a bulk sample. For the method with spherical harmonics model cite Liebi et. al. Nature 527, 349 (2015) and Liebi et. al. Acta Cryst. A74, 12 (2018). For the IRTT method cite Gao et. al. Acta Cryst. A75, 223 (2019). Uses the base and scanning SAXS packages above.
Tomography package
Beamline codes for alignment of projections and reconstruction for tomography and laminography. Projection alignment methods by 3D consistency with deep sub-pixel accuracy are included. If used cite Odstrčil et al. Opt. Express 25, 36637 (2019). Uses the base package above.
PtychoShelves
A versatile high-level framework for high-performance analysis of ptychographic data. This package is used for preparation and reconstruction of data from cSAXS as well as other beamlines and synchrotrons. It allows for modular use of several data inputs and reconstruction algorithms. Includes Matlab prototype codes as well as highly optimized C code both for CPU and GPU. If used cite Wakonig et al. J. Appl. Cryst. 53, 574-586 (2020). Uses the base package above.
Other open source codes
LSQ-ML - Iterative least-squares maximum-likelihood ptychography solver
This source code provides an implementation of the LSQ-ML method described in Opt. Express 26, 3108 (2018) in a simple Matlab-based framework. The reference implementation includes an artificial data generator and implementations of the ePIE, ePIE-OPRP and difference-map codes that were used in the article. Note that this is not the cSAXS ptychography package (which is planned to be published here at a later date), but rather a simplified version intended to showcase the LSQ-ML algorithm.
The code is using Matlab-based GPU acceleration, however seamless fallback CPU option is available as well.
Alignment methods for nanotomography with deep sub-pixel accuracy
This repository contains codes for alignment of projections and reconstruction for tomography and laminography. Alignment methods by 3D consistency with deep sub-pixel accuracy are included. Additionally we include artificial, i.e. simulated, and experimental examples. If used cite Odstrčil et. al. Opt. Express 25, 36637 (2019).